Data Analysis (Module 2, p. 2/2)
Appendix Module 2
- Full correlation matrix (by all stations) here…
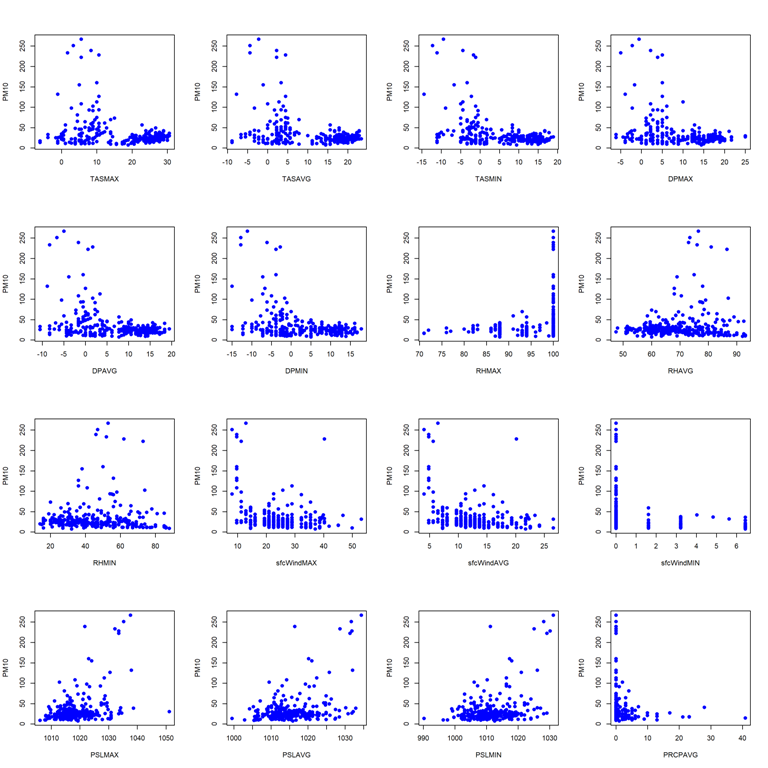
- Pairwise scatterplots for station 9572
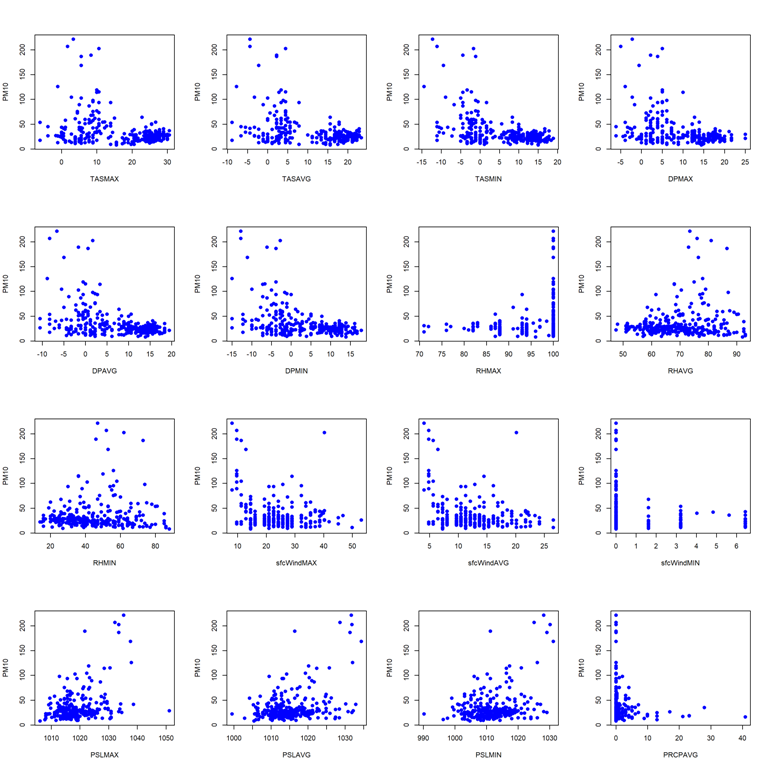
- Pairwise scatterplots for station 9616
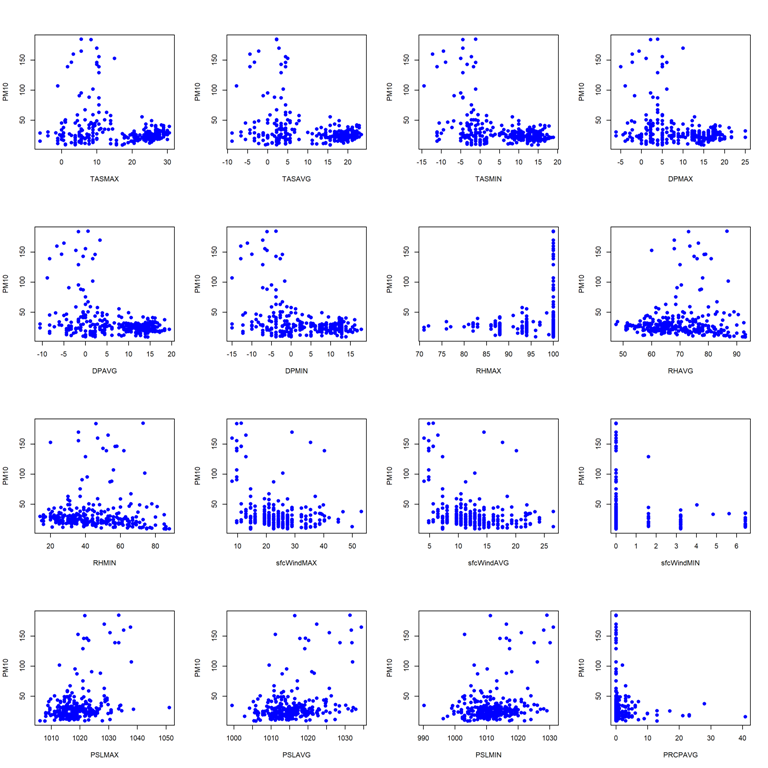
- Pairwise scatterplots for station 9642
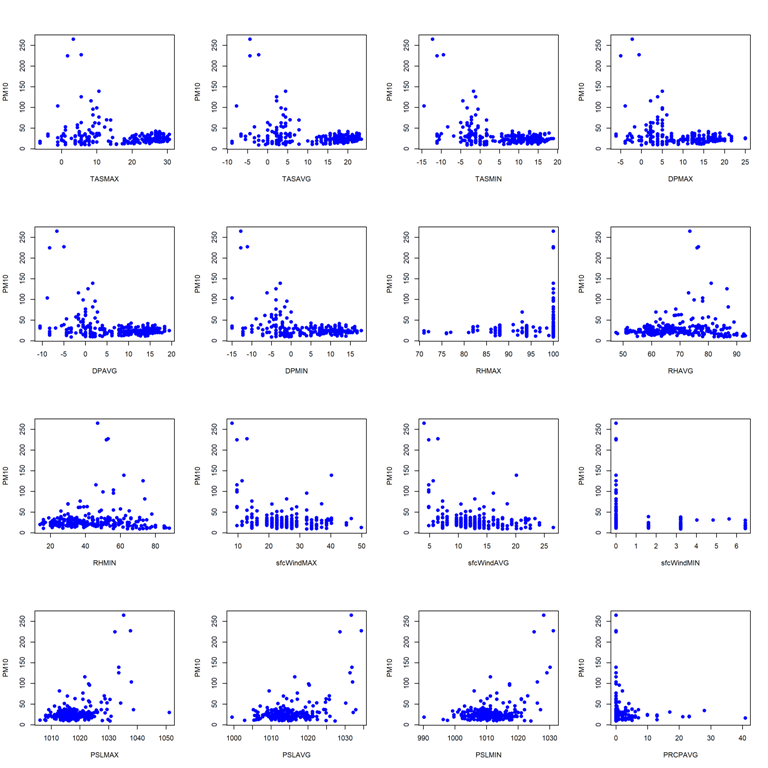
- Pairwise scatterplots for station 60881
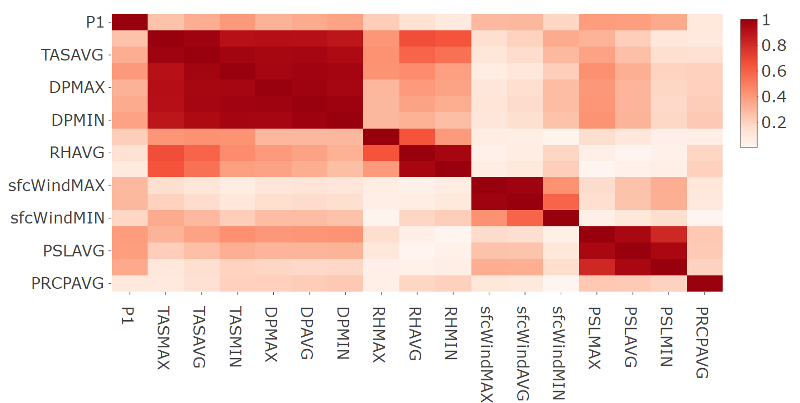
- Plot of the correlation matrix in absolute terms for station 9572
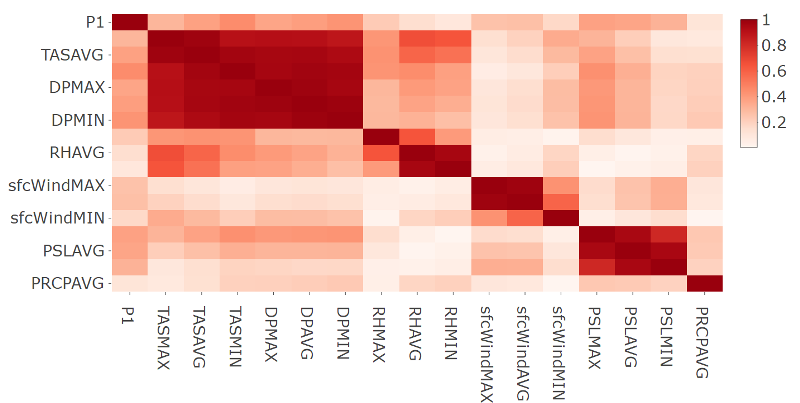
- Plot of the correlation matrix in absolute terms for station 9616
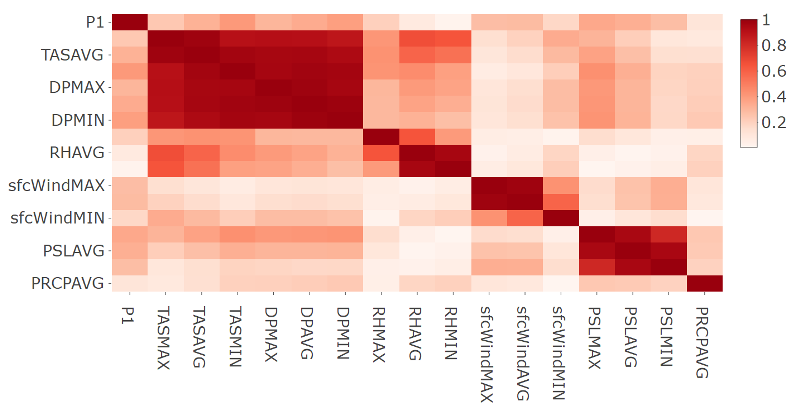
- Plot of the correlation matrix in absolute terms for station 9642
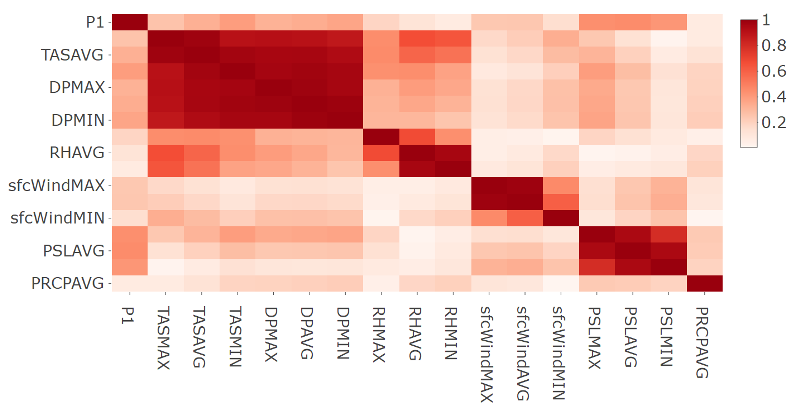
- Plot of the correlation matrix in absolute terms for station 60881
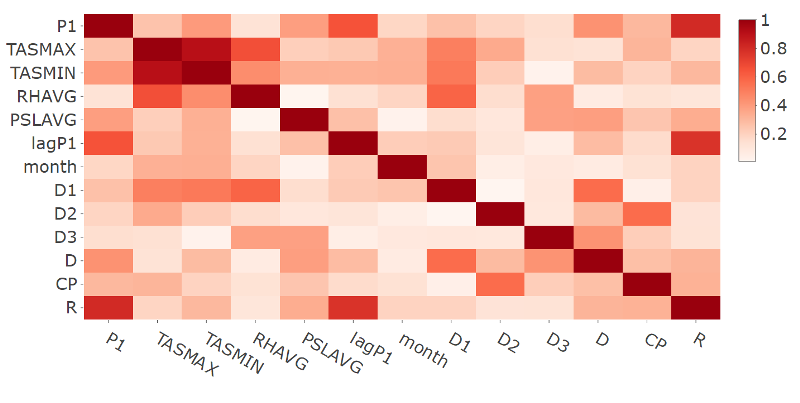
- Plot of the correlation matrix in absolute terms for the response and selected features variables for station 9572
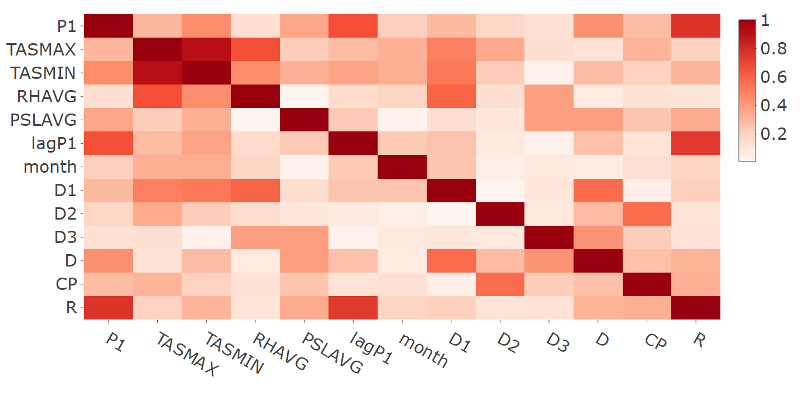
- Plot of the correlation matrix in absolute terms for the response and selected features variables for station 9616
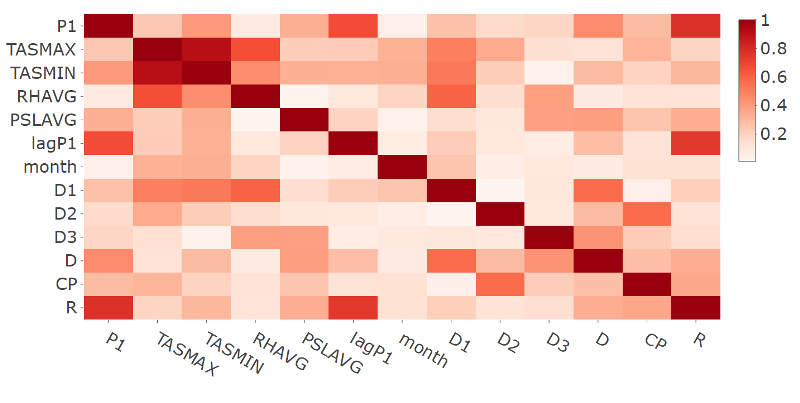
- Plot of the correlation matrix in absolute terms for the response and selected features variables for station 9642
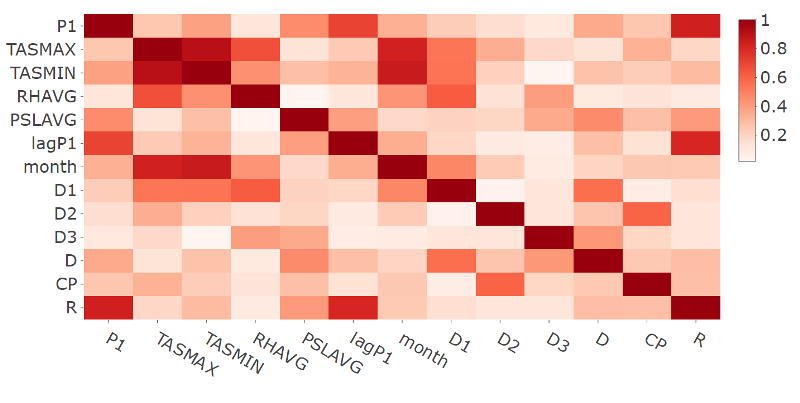
- Plot of the correlation matrix in absolute terms for the response and selected features variables for station 60881
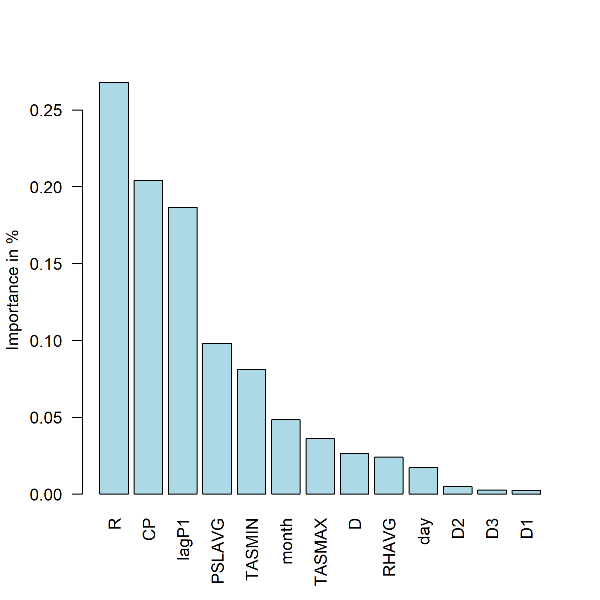
- Plot: Summary of feature importance expressed as percentage of the cumulative decrease in the optimized loss function. Station 9572
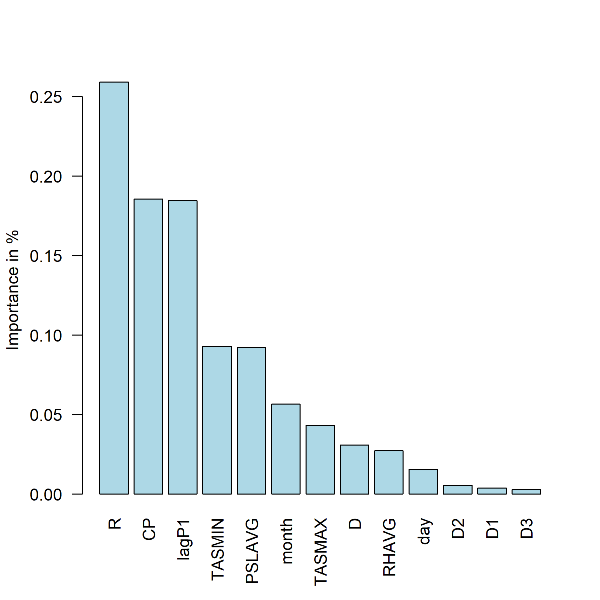
- Plot: Summary of feature importance expressed as percentage of the cumulative decrease in the optimized loss function. Station 9616
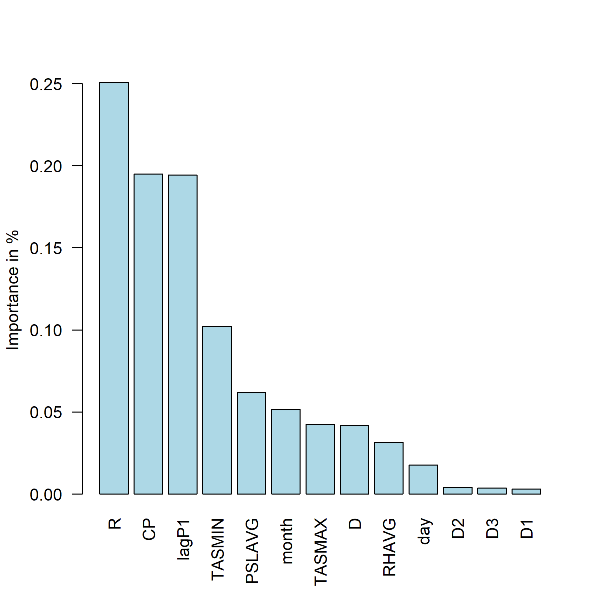
- Plot: Summary of feature importance expressed as percentage of the cumulative decrease in the optimized loss function. Station 9642
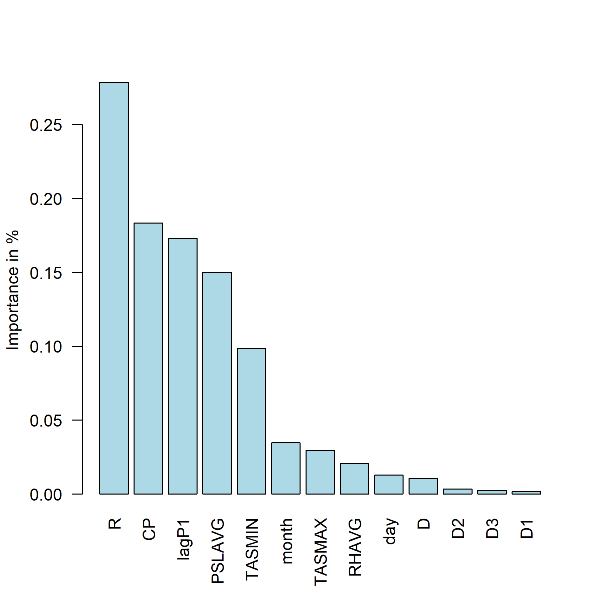
- Plot: Summary of feature importance expressed as percentage of the cumulative decrease in the optimized loss function. Station 60881
Code
Download the R code here… Download the data here…
Set up environment
rm(list=ls())
gc()
Set the path of the folder with data
setwd("/data")
Import data on EEU measurements for 2017 and 2018
eeu=list.files(path="/data",pattern="BG*")
ddeu=lapply(eeu,read.csv,na.string=c("","NA"," "), stringsAsFactors = F, fileEncoding="UTF-16LE")
Name data sets by stations
for (i in 1:length(eeu)){
eeu[i]=gsub("BG_5_","st", eeu[i])
eeu[i]=gsub("_timeseries.csv","", eeu[i])
names(ddeu)[i]=eeu[i]
}
rm(eeu,i)
Select only the observations with averaging time == “hour”
for (i in 1:length(ddeu)){
ddeu[[i]]=ddeu[[i]][ddeu[[i]]$AveragingTime=="hour",]
}
count = 0
i = 1
while(0 == 0){
if (count == 1){
i = i - 1
}
if(i > length(ddeu)){
break
}
count = 0
if(dim(ddeu[[i]])[1] == 0){
ddeu[[i]] = NULL
count = 1
}
i = i + 1
}
rm(count,i)
Check variables’ class of ddeu
sapply(lapply(ddeu,"[", ,"DatetimeEnd"),class)
sapply(lapply(ddeu,"[", ,"Concentration"),class)
Fix the class of the time variable
if(!require(lubridate)){
install.packages("lubridate")
library(lubridate)
}
for (i in 1:length(ddeu)){
ddeu[[i]]=ddeu[[i]][,c("DatetimeEnd","Concentration")]
ddeu[[i]]$DatetimeEnd=ymd_hms(ddeu[[i]]$DatetimeEnd, tz="Europe/Athens")
colnames(ddeu[[i]])=c("time","P1eu")
}
sapply(lapply(ddeu,"[", ,"time"),class)
Make a time vecor with hourly sampling rate
teu=list()
for (i in 1:length(ddeu)){
teu[[i]]=as.data.frame(seq.POSIXt(from=min(ddeu[[i]]$time),to=max(ddeu[[i]]$time), by="hour"))
colnames(teu[[i]])[1]="time"
}
Make a list of time series on official P10 concentration for every station
if(!require(dplyr)){
install.packages("dplyr")
library(dplyr)
}
if(!require(devtools)){
install.packages("devtools")
library(devtools)
}
for (i in 1:length(ddeu)){
teu[[i]]=left_join(teu[[i]],ddeu[[i]],by="time")
}
names(teu)=names(ddeu)
Bind datasets 2017 and 2018 for each of the official points
teu$st9421=bind_rows(teu$st9421_2017,teu$st9421_2018)
teu$st9572=bind_rows(teu$st9572_2017,teu$st9572_2018)
teu$st9616=bind_rows(teu$st9616_2017,teu$st9616_2018)
teu$st9642=bind_rows(teu$st9642_2017,teu$st9642_2018)
teu$st60881=teu$st60881_2018
teu=teu[c("st9421", "st9572","st9616","st9642","st60881")]
for (i in 1:length(teu)){
colnames(teu[[i]])[2]="P1"}
rm(i,ddeu)
Check for duplicates
sapply(teu,dim)[1,]
sum(duplicated(teu[[1]]$time)) #0
sum(duplicated(teu[[2]]$time)) #0
sum(duplicated(teu[[3]]$time)) #0
sum(duplicated(teu[[4]]$time)) #0
sum(duplicated(teu[[5]]$time)) #0
Interpolate missing values for P1eu
if(!require(imputeTS)){
install.packages("imputeTS")
library(imputeTS)
}
Check the numer of missing obs
sapply(sapply(teu,is.na),sum)
sapply(sapply(teu,is.na),sum)/sapply(teu,dim)[1,]
Apply linear interpolation
for (i in 1:length(teu)){
teu[[i]][,2]=na.interpolation(teu[[i]][,2], option="linear")
}
rm(i)
Aggregate official measuremnets on daily basis
Extract date from the time variable
for (i in 1:length(teu)){
teu[[i]]$date=date(teu[[i]]$time)
}
Aggregate data on daily basis
aeu=list()
for (i in 1:length(teu)){
aeu[[i]]=teu[[i]] %>%
group_by(date) %>%
summarise(P1=mean(P1))
}
names(aeu)=names(teu)
#rm(teu,i)
Import data on weather metrics
Import data on weather
setwd("/")
ww=read.csv("lbsf_20120101-20180917_IP.csv",na.string=c("","NA"," ",-9999), stringsAsFactors = F)
Get a date vector
ww$date=make_date(year=ww$year,month=ww$Month,day=ww$day)
ww=ww[,c(23,1:22)]
Check for poorly populated variables
colSums(is.na(ww))
ww$date[is.na(ww$VISIB)==T]
Entire period for PRCPMAX and PRCPMIN is missing Last two months (Aug and Sep 2018) for VISIB are missing Remove this variables from the sample
ww=ww[,!names(ww) %in% c("PRCPMAX","PRCPMIN","VISIB","year","Month","day")]
Merge data on P10 and weather
for (i in 1:length(aeu)){
aeu[[i]]=left_join(aeu[[i]],ww,by="date")
aeu[[i]]$day=wday(aeu[[i]]$date,label=T)
}
Handle missing values
sapply(sapply(aeu,is.na),colSums)
for (i in 1:length(aeu)){
aeu[[i]][,3:18]=sapply(aeu[[i]][,3:18],na.interpolation,option="linear")
}
rm(ww,i)
Look at the data first
Construct correlation matrix
cc=list()
for (i in 1:length(aeu)){
cc[[i]]=cor(aeu[[i]][,2:18])
}
names(cc)=names(aeu)
Visualize correlation matrix
if(!require(plotly)){
install.packages("plotly")
library(plotly)
}
if(!require(RColorBrewer)){
install.packages("RColorBrewer")
library(RColorBrewer)
}
i=1
Change i=1,2,3,4,5 so as to get plot for each station
plot_ly(x=rownames(cc[[i]])[1:nrow(cc[[i]])], y=colnames(cc[[i]])[nrow(cc[[i]]):1],z=abs(cc[[i]][nrow(cc[[i]]):1,1:nrow(cc[[i]])]),type="heatmap",colors=brewer.pal(8,"Reds"))
Stationarity tests
if(!require(tseries)){
install.packages("tseries")
library(tseries)
}
aux=list()
for (i in 1:length(aeu)){
a=adf.test(aeu[[i]]$P1)
aux[[i]]=c(a$statistic,a$p.value)
rm(a)
}
aux=as.data.frame(aux)
colnames(aux)=names(aeu)
rownames(aux)=c("ADF statistics", "p-value")
Feature engineering
eu=list()
for (i in 1:length(aeu)){
eu[[i]]=aeu[[i]][,c("P1","TASMAX","TASMIN","RHAVG","PSLAVG","day")]
eu[[i]]$lagP1=dplyr::lag(eu[[i]]$P1,1)
eu[[i]]$month=month(aeu[[i]]$date)
eu[[i]]$D1=ifelse(aeu[[i]]$RHMAX==100,1,0)
eu[[i]]$D2=ifelse(aeu[[i]]$sfcWindMIN==0,1,0)
eu[[i]]$D3=ifelse(aeu[[i]]$PRCPAVG==0,1,0)
eu[[i]]$D=eu[[i]]$D1*eu[[i]]$D2*eu[[i]]$D3
eu[[i]]$CP=aeu[[i]]$sfcWindAVG*(dplyr::lag(aeu[[i]]$sfcWindAVG,1))
eu[[i]]$R=eu[[i]]$lagP1/eu[[i]]$CP
eu[[i]]=eu[[i]][-1,]
}
names(eu)=names(aeu)
Construct correlation matrix
cc=list()
for (i in 1:length(eu)){
cc[[i]]=cor(eu[[i]][,!names(eu[[i]]) %in% "day"])
}
names(cc)=names(eu)
Visualize correlation matrix
i=1 ``` ```R Change i=1,2,3,4,5 so as to get plot for each station
plot_ly(x=rownames(cc[[i]])[1:nrow(cc[[i]])], y=colnames(cc[[i]])[nrow(cc[[i]]):1],z=abs(cc[[i]][nrow(cc[[i]]):1,1:nrow(cc[[i]])]),type="heatmap",colors=brewer.pal(8,"Reds"))
Random forest
mean absolute percentage error
MAE=list()
root mean squared error
RMSE=list()
feature importance
FI=list()
for (i in 1:length(eu)){
FI[[i]]=data.frame(matrix(NA,ncol=1,nrow=13))
FI[[i]][,1]=as.character(names(eu[[i]])[-1])
colnames(FI[[i]])="fname"
}
names(FI)=names(eu)
for (t in 1:100){
Split data into training and test set
set.seed(t)
train=list()
for (i in 1:length(eu)){
train[[i]]=sample(nrow(eu[[i]]),round(nrow(eu[[i]])/2))
}
Model estimation & feature importance
if(!require(randomForest)){
install.packages("randomForest")
library(randomForest)
}
eq=list()
for (i in 1:length(eu)){
eq[[i]]=randomForest(P1~.,data=eu[[i]][train[[i]],])
a=data.frame(fname=as.character(rownames(eq[[i]]$importance)), imp=as.numeric(eq[[i]]$importance))
colnames(a)[2]=paste("imp",t,sep="")
FI[[i]]=left_join(FI[[i]],a,by="fname")
rm(mae,rmse,a)
}
Calculate out-of-sample accuracy
mae=data.frame(matrix(NA,nrow=length(eu),ncol=3))
rownames(mae)=names(eu)
colnames(mae)=c("M1","M2","M3")
rmse=data.frame(matrix(NA,nrow=length(eu),ncol=3))
rownames(rmse)=names(eu)
colnames(rmse)=c("M1","M2","M3")
for (i in 1:length(eu)){
f=predict(eq[[i]],newdata=eu[[i]][-train[[i]],])
rmse[i,"M1"]=sqrt(mean((eu[[i]]$P1[-train[[i]]]-mean(eu[[i]]$P1[-train[[i]]]))^2))
rmse[i,"M2"]=sqrt(mean((eu[[i]]$P1[-train[[i]]]-eu[[i]]$lagP1[-train[[i]]])^2))
rmse[i,"M3"]=sqrt(mean((eu[[i]]$P1[-train[[i]]]-f)^2))
mae[i,"M1"]=mean(abs((eu[[i]]$P1[-train[[i]]]-mean(eu[[i]]$P1[-train[[i]]]))/eu[[i]]$P1[-train[[i]]]))
mae[i,"M2"]=mean(abs((eu[[i]]$P1[-train[[i]]]-eu[[i]]$lagP1[-train[[i]]])/eu[[i]]$P1[-train[[i]]]))
mae[i,"M3"]=mean(abs((eu[[i]]$P1[-train[[i]]]-f)/eu[[i]]$P1[-train[[i]]]))
}
MAE[[t]]=mae
RMSE[[t]]=rmse
}
rm(i,t)
MAE=as.data.frame(MAE)
RMSE=as.data.frame(RMSE)
mae=data.frame(matrix(NA,nrow=length(eu),ncol=3))
rownames(mae)=names(eu)
colnames(mae)=c("M1","M2","M3")
rmse=data.frame(matrix(NA,nrow=length(eu),ncol=3))
rownames(rmse)=names(eu)
colnames(rmse)=c("M1","M2","M3")
Devrie rmse and mae
for (i in 1:length(eu)){
mae[i,"M1"]=mean(as.numeric(MAE[i,grep("^M1",names(MAE))]))
mae[i,"M2"]=mean(as.numeric(MAE[i,grep("^M2",names(MAE))]))
mae[i,"M3"]=mean(as.numeric(MAE[i,grep("^M3",names(MAE))]))
rmse[i,"M1"]=mean(as.numeric(RMSE[i,grep("^M1",names(RMSE))]))
rmse[i,"M2"]=mean(as.numeric(RMSE[i,grep("^M2",names(RMSE))]))
rmse[i,"M3"]=mean(as.numeric(RMSE[i,grep("^M3",names(RMSE))]))
}
Derive importance
fi=list()
for (i in 1:length(FI)){
fi[[i]]=data.frame(fname=FI[[i]]$fname)
fi[[i]]$fimp=apply(FI[[i]][,2:101],1,mean)
fi[[i]]$pfimp=fi[[i]]$fimp/sum(fi[[i]]$fimp)
rownames(fi[[i]])=FI[[i]]$fname
}
i=5
Change i=1,2,3,4,5 so as to get plot for each station
barplot(fi[[i]]$pfimp[order(fi[[i]]$pfimp,decreasing = T)], names.arg = fi[[i]]$fname[order(fi[[i]]$pfimp,decreasing = T)],col="light blue", ylab="Importance in %",las=2)
Get predition for P10 by stations
NB: This prediction assumes we have weather forecast Use the following code:
f=list()
for (i in 1:length(eu)){
f[[i]]=predict(eq[[i]],newdata=[\#inseart here new data])
}
[Acknowledgment] [Introduction] [Methodology] [Bias correction] [Analysis] [Features] [Prediction] [Summary]